Learning Gabor filters with ICA and scikit-learn
My colleague Hannes works in deep learning and started on a new feature extraction method this week.
As with all feature extraction algorithms, it was obviously of utmost importance to be able to learn Gabor filters.
Inspired by his work and Natural Image Statistics, a great book on the topic of feature extraction from images, I wanted to see how hard it is to learn Gabor filters with my beloved scikit-learn.
I chose independent component analysis, since this is discusses to some depth in the book.
Luckily mldata had some image patches that I could use for the task.
Here goes:
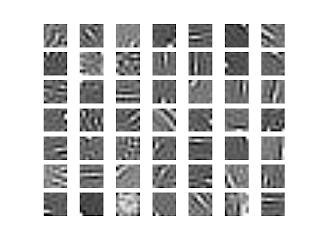
And the result (takes ~40sec):
The data is somewhat normalized already, which makes the job easier. With larger patches, it does not look as clean. I guess it is because there is not enough data. It would be possible to use PCA to get rid of "high frequencies" but at this small patch size, it did not seem to matter.
Edit:
As Andrej suggested, here the k-means filters on whitened data:
For reference, the PCA whitening filters:
which are the expected wavelets.
You can find the updated gist here.
There is also a nice example of learning similar filters from lena using scikits-image.
As with all feature extraction algorithms, it was obviously of utmost importance to be able to learn Gabor filters.
Inspired by his work and Natural Image Statistics, a great book on the topic of feature extraction from images, I wanted to see how hard it is to learn Gabor filters with my beloved scikit-learn.
I chose independent component analysis, since this is discusses to some depth in the book.
Luckily mldata had some image patches that I could use for the task.
Here goes:
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import fetch_mldata
from sklearn.decomposition import FastICA
# fetch natural image patches
image_patches = fetch_mldata("natural scenes data")
X = image_patches.data
# 1000 patches a 32x32
# not so much data, reshape to 16000 patches a 8x8
X = X.reshape(1000, 4, 8, 4, 8)
X = np.rollaxis(X, 3, 2).reshape(-1, 8 * 8)
# perform ICA
ica = FastICA(n_components=49)
ica.fit(X)
filters = ica.unmixing_matrix_
# plot filters
plt.figure()
for i, f in enumerate(filters):
plt.subplot(7, 7, i + 1)
plt.imshow(f.reshape(8, 8), cmap="gray")
plt.axis("off")
plt.show()
And the result (takes ~40sec):
The data is somewhat normalized already, which makes the job easier. With larger patches, it does not look as clean. I guess it is because there is not enough data. It would be possible to use PCA to get rid of "high frequencies" but at this small patch size, it did not seem to matter.
Edit:
As Andrej suggested, here the k-means filters on whitened data:
For reference, the PCA whitening filters:
which are the expected wavelets.
You can find the updated gist here.
There is also a nice example of learning similar filters from lena using scikits-image.






If Gabors is what you're after, scikits has had k-means for a very long time!
ReplyDeleteThat's true. It has had ICA for a long time, too. Just thought it would be a nice thing to play around with. But you are right, doing k-means would also be interesting.
ReplyDeleteDictionary learning with an l1 penalty will also give you Gabors. Anyhow, k-means, ICA and sparse dictionary learning all seek similar patterns in the data: sparse or super Gaussian loadings. I wonder if NMF would give you Gabors....
ReplyDeleteNMF can not give you Gabor filters, as you can not produce a negative part. If you do parse NMF and have a matrix that is restricted to be positive and subtract one that is also restricted to be positive, the two can combine to Gabor filters.
ReplyDeleteThis is discussed in the book I mentioned.
I thought so, but I've been surprised before. I didn't expect to get real Gabors, but I was wondering if you would get the positive part of Gabors.
ReplyDeletehow can one apply these filters on a image? Is it by Convolution?
ReplyDeletewhich scikit version are you using? In mine, it throws error as FastICA does not have unmixing_matrix_ object
ReplyDeleteThis has been a while. In newer versions, it's called ``components_``.
Delete