Simplistic Minimum Spanning Tree in Numpy [update]
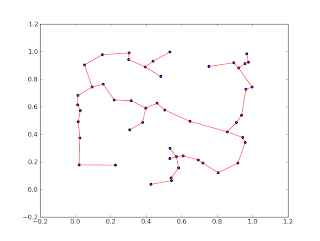
I started working with spanning trees for euclidean distance graphs today. The first think I obviously needed to do was compute the spanning tree. There are MST algorithms in Python, for example in pygraph and networkx . These use their native graph formats, though, which would have meant I'd have to construct a graph from my point set. I didn't see a way on how to do this and set the edge weights without iterating over all edges. That would probably take longer than the computation of the MST, so I decided to do my own small implementation using numpy. This is an instantiation of Prim's algorithm based on numpy matrices. The input is a dense matrix of distances, the output a list of edges. It is not as pretty as I would have hoped but still reasonably short. If any one has suggestions how to make this prettier, I'd love that. [edit] Using line_profiler I had a quick look a the code and made some minor improvements. It's now significantly faster than net...